CustardPy#
CustardPy is a 3D genome analysis tool written in Python3 and available using the Docker system. It is primarily designed for multi-sample Hi-C analysis (e.g., comparing depletion effects across multiple proteins) and offers various functions for analysis and visualization. Since the CustardPy docker image includes various existing tools in addition to the core component (see Installation), users can perform a variety of 3D genome analyses without installing them individually.
A common problem in Hi-C analysis is the strict requirement of specific input formats. Many tools require input data to be in a specific format, and consequently, their use is hindered if the data under investigation does not conform to these specifications. Since CustardPy covers the processing of Hi-C data from FASTQ and uses the generated data for subsequent analysis, users can avoid potential format incompatibilities.
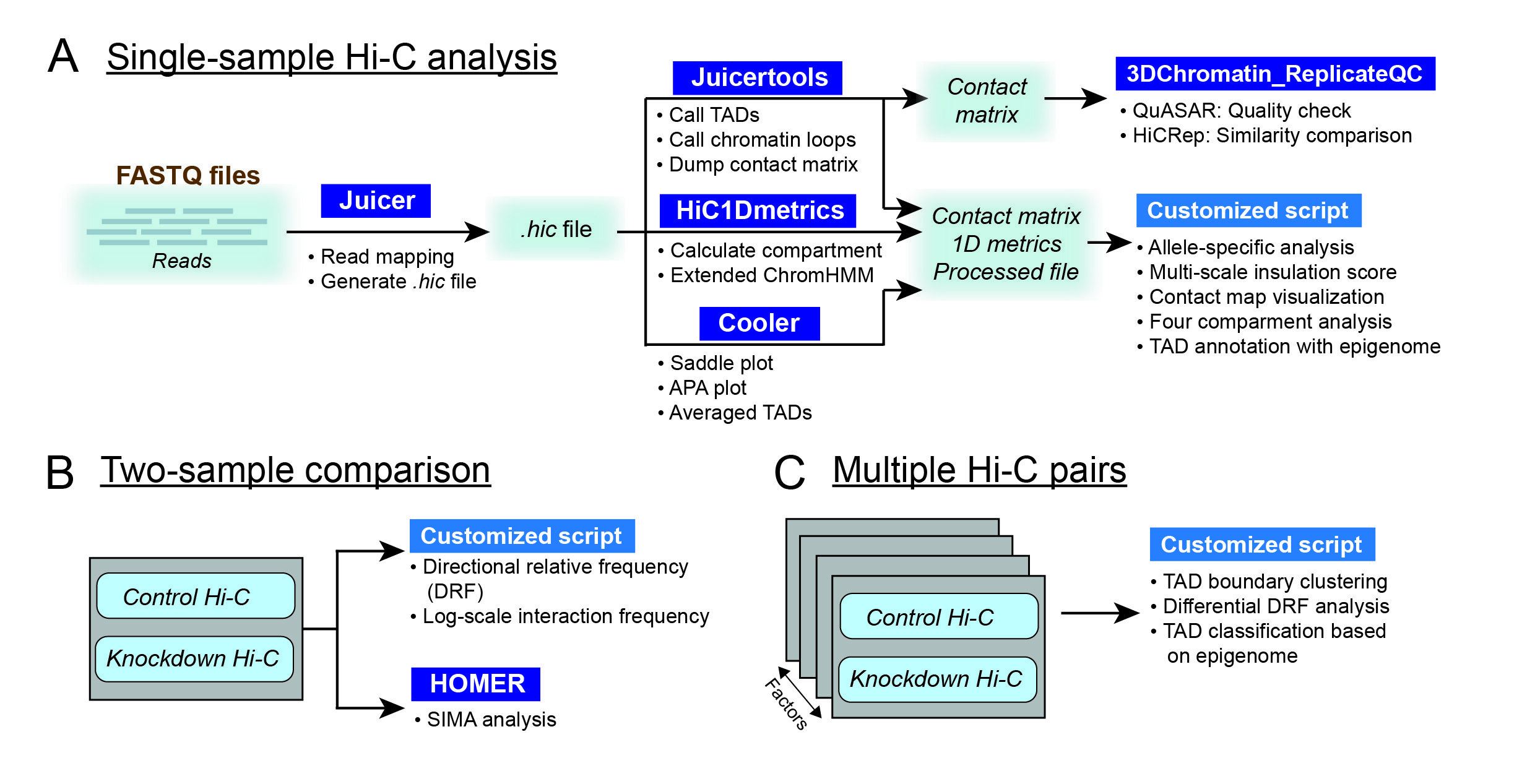
Fig. 1 The workflow of CustardPy for a single sample (A), two samples (B) and multiple sample comparison (C).#
Contents:#
Citation:#
Nakato R, Sakata T, Wang J, Nagai LAE, Nagaoka Y, Oba GM, Bando M, Shirahige K, Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors, Nature Communications, 2023. doi: 10.1038/s41467-023-41316-4
